|
Classical attenuators monitor the ability of the ribosome to translate a short leader peptide. Attenuators are sensory RNAs as they respond directly to environmental signals by toggling between alternative secondary structures either favoring or preventing expression of downstream genes ( Naville and Gautheret, 2010 Mellin and Cossart, 2015). RNA attenuators are part of the mRNA that they regulate and therefore act in cis. Regulatory RNAs include two major classes, which are RNA attenuators and small RNAs (sRNAs) ( Henkin, 2008). 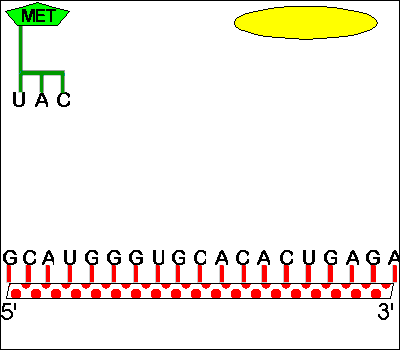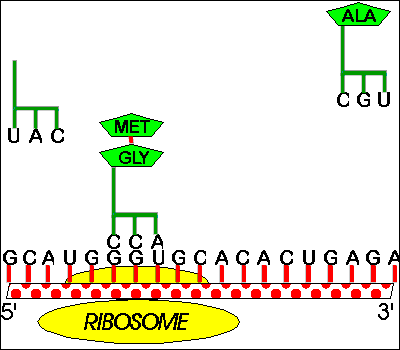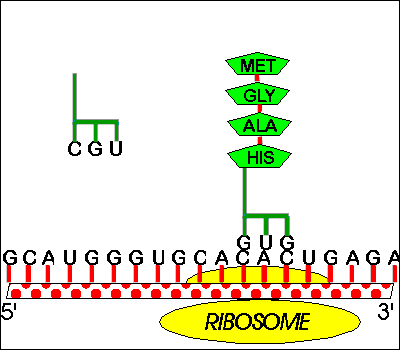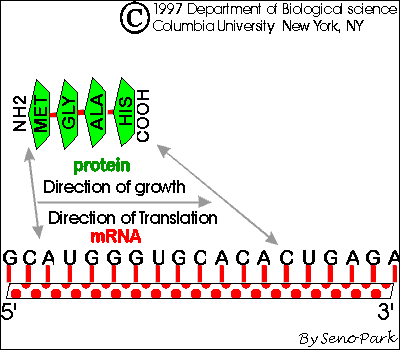
These RNAs are often referred to as non-coding RNAs (ncRNAs) as they usually operate on their own without the need for being translated ( Repoila and Darfeuille, 2009). Regulatory RNAs in bacteria comprise a heterogeneous group of molecules that act by various mechanisms to modulate cellular processes in response to cognate stimuli. Evidence is accumulating that regulatory RNAs are also important players for the bacterial response and resistance to antibiotics, making these molecules promising targets for antimicrobial chemotherapy. In bacteria, regulatory RNAs act at the post-transcriptional level to control bacterial physiology, development, and virulence ( Oliva et al., 2015). Traditionally, efforts to find novel treatment options have focussed on bacterial proteins as drug targets, whereas exploiting regulatory RNA elements was only considered of late. The emergence and spread of resistance to antibiotics represent a major threat for human health and urgently call for novel antimicrobial compounds and therapies. Approaches to target RNAs involved in intrinsic antibiotic resistance or virulence for chemotherapy will be discussed. In this review, we address how regulatory RNAs respond to and establish resistance to antibiotics in bacteria. Drugs acting on sRNAs are considered for combined therapies to treat infections. 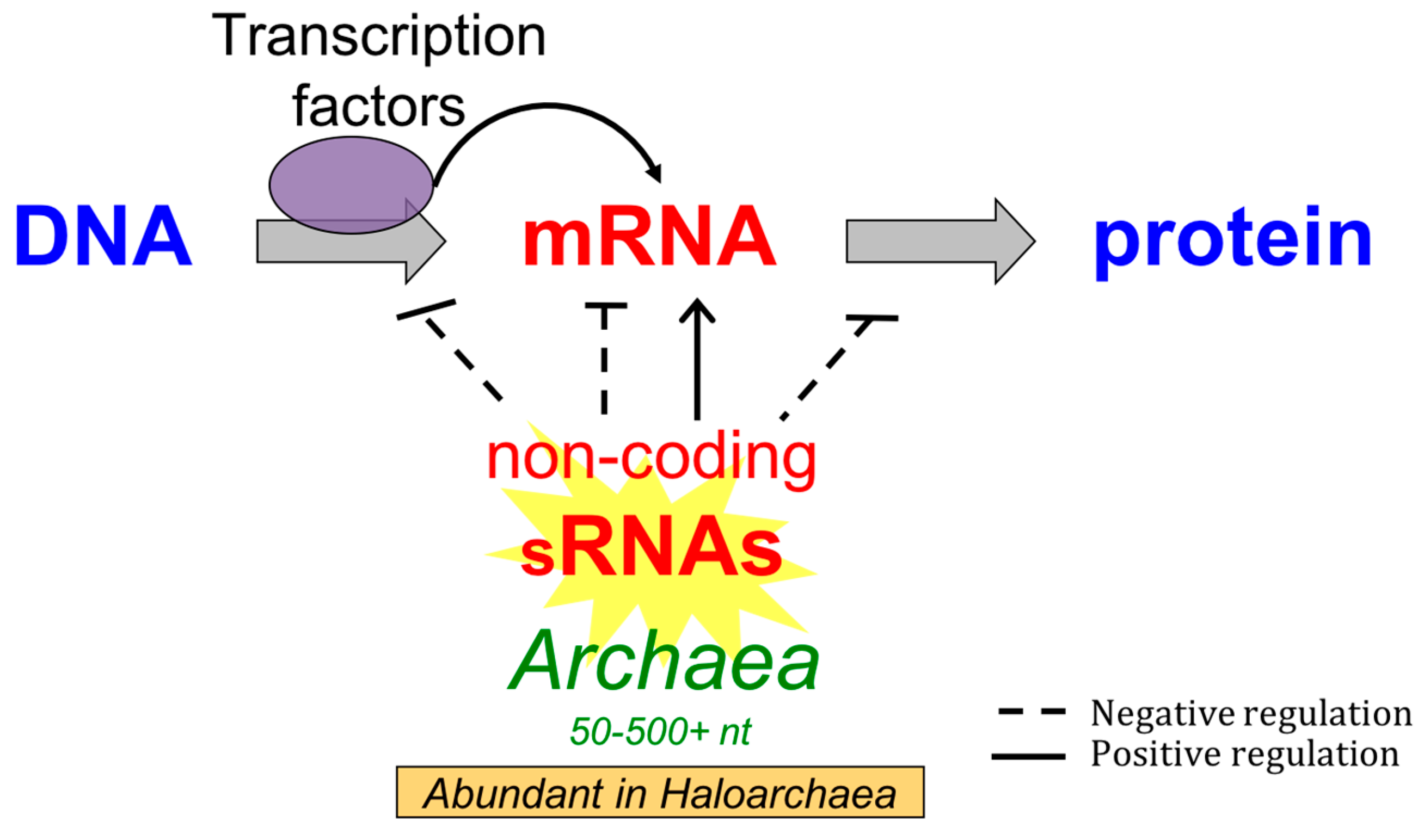
Indeed, several compounds have been developed that kill bacteria by mimicking ligands for riboswitches controlling essential genes, demonstrating that regulatory RNA elements are druggable targets. In consequence, regulatory RNAs including sRNAs and their protein interaction partners such as Hfq may prove useful as targets for antimicrobial chemotherapy. Each antibiotic generates a unique sRNA profile possibly causing downstream effects that may help to overcome the antibiotic challenge. Bacteria respond with extensive changes of their sRNA repertoire to antibiotics. Trans-encoded small RNAs (sRNAs) modulate antibiotic tolerance by base-pairing with mRNAs encoding functions important for resistance such as metabolic enzymes, drug efflux pumps, or transport proteins. RNA attenuator elements in mRNA leader regions couple expression of resistance genes to the presence of the cognate antibiotic. Lately, the contribution of non-coding RNAs to antibiotic resistance and their potential value as drug targets became evident. 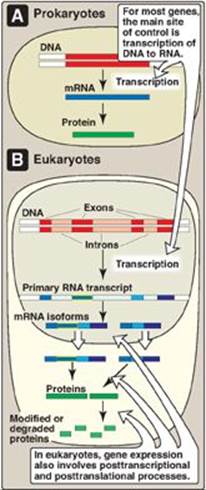
The emergence of antibiotic resistance mechanisms among bacterial pathogens increases the demand for novel treatment strategies. Perutz Laboratories, University of Vienna, Vienna, Austria 2Department of Microbiology, Immunobiology and Genetics, Max F.1Department of Molecular Infection Biology, Helmholtz Centre for Infection Research, Braunschweig, Germany.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
